Featured Faculty
Professor Xiangyun Qiu

Dr. Qiu studies molecular biophysics using advanced x-ray and neutron scattering and spectroscopic approaches
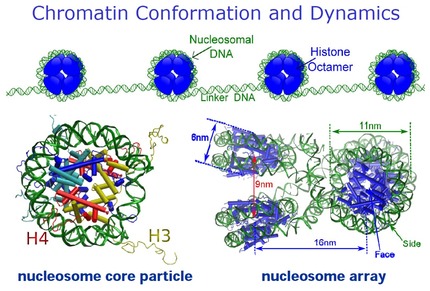
It is a fundamental question in biology to understand chromatin
structure and its roles in gene regulation. The objective of our
research is to gain a detailed and mechanistic understanding of the
conformation and dynamics of chromatin modulated by solution conditions
(ions and molecular crowding) and genetic and epigenetic variations
(nucleosomal and linker DNA and histone). Eukaryotic cells organize
their genome in the form of chromatin with the nucleosome as its
elemental unit. While chromatin is closely involved in DNA-directed
processes such as transcription, replication, recombination, and repair,
whether and how the behavior of chromatin itself, the “substrate”
therein, regulates such processes remains an open question. One critical
barrier to answer this question is the lack of quantitative knowledge
of the conformation and dynamics of chromatin beyond the nucleosome. To
this end, we take a multi-length-scale approach that integrates solution
small angle x-ray and neutron scattering (SAXS/SANS), Förster resonance
energy transfer (FRET), and buffer equilibrium atomic emission
spectroscopy (BE-AES) to probe global, local, and charge-specific
structures, respectively. Well-defined systems of recombinant nucleosome
arrays with increased number of repeats and distinctive sequence
features will be studied in vitro. The goal of these bottom-up studies
is to develop quantitative models of chromatin structures guided by
systematic measurements.